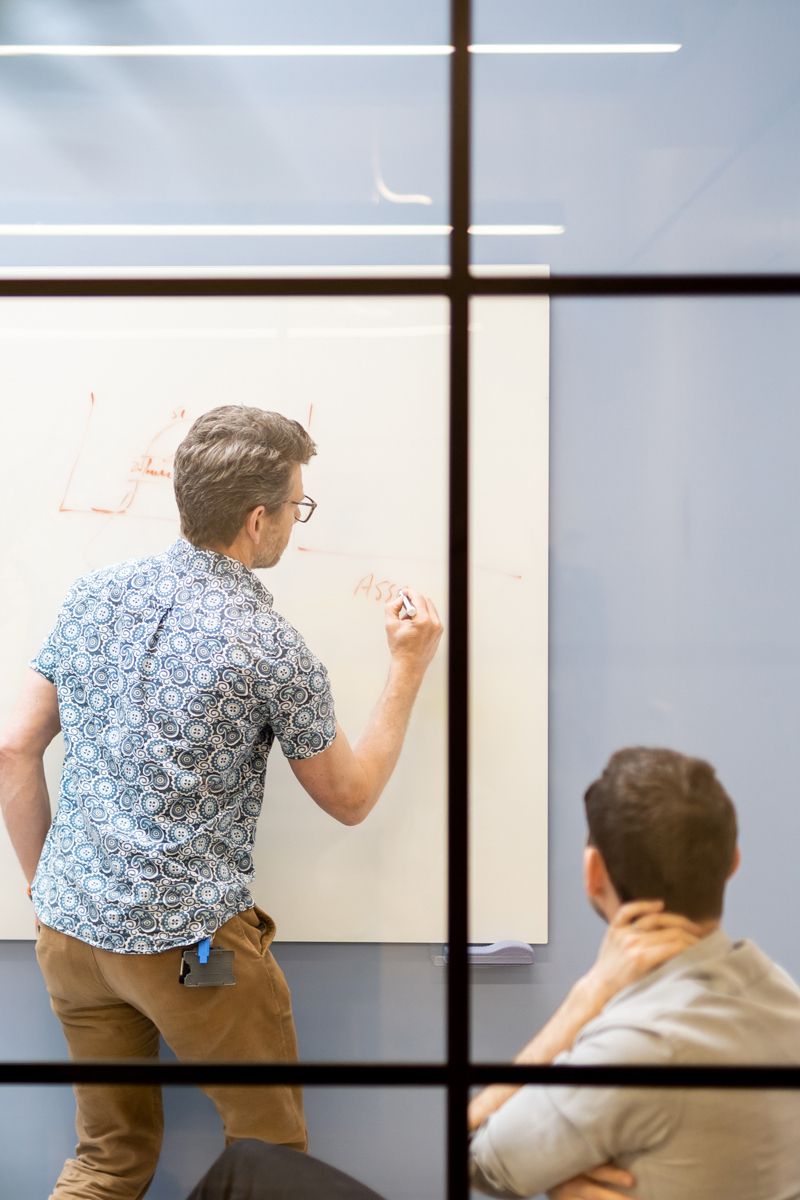
A versatile, scalable and robust AI-enabled drug discovery platform built with expert scientists, leveraging multi-modal data foundations.
End-to-end Drug Discovery
The Benevolent PlatformTM integrates technology, processes, and humans for faster and higher-quality R&D success
- Technology tailored for discovery with:
- Unique data foundations from multiple data types curated and purpose-built for drug discovery
- Explainable AI models that enable scientists to see rationale for predictions
- Applications across therapeutic areas and modalities
- Precision workflows merge the strengths of AI & data analytics with insights derived from scientists’ strategic oversight and intuition
- R&D Experts employ the tools to ensure that deep domain expertise guides every phase of development
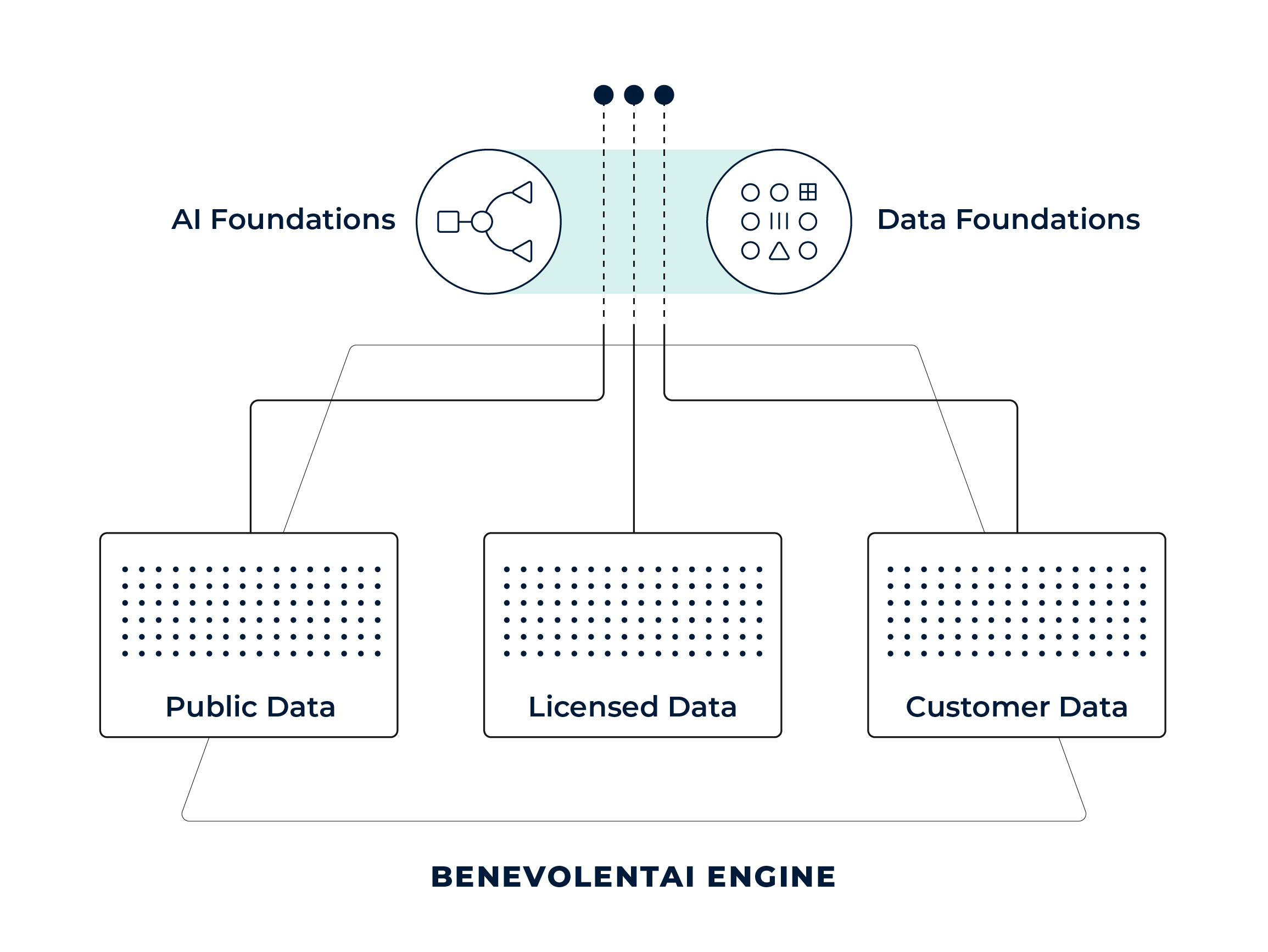
AI FOUNDATIONS
BenAI Engine
BenAI Engine is the core foundation of our technology that turns fragmented scientific data into actionable insights.
It is designed to ingest, organise, and leverage biomedical data for a nuanced understanding of disease biology.
BENEFITS
Proven to enhance drug discovery
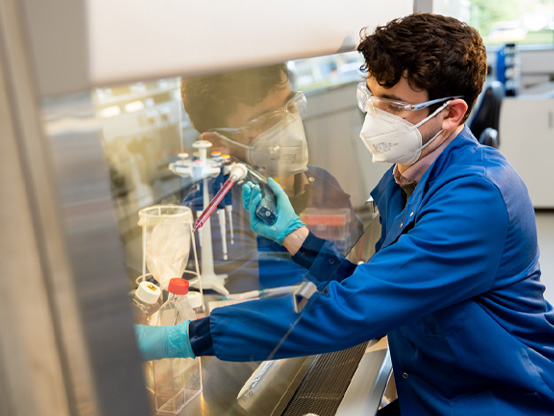

WET LABS
In-house wet labs reduce development timelines
Our fully equipped laboratories in Cambridge (UK), and drug discovery team can rapidly progress programmes leveraging our cutting-edge technologies such as CRISPR, RNA-seq and human iPSCs.
PARTNERING
We partner with leading pharmaceutical and biotech companies to help them unlock biological insights and tackle complex therapeutic challenges.
