There are moral, scientific and economic reasons for having a more inclusive patient demographic represented in precision medicine.
Precision medicine promises to provide better matches between treatments and groups of patients who have a disease and who show a set of specific traits. Better matches help ensure that a drug would be given to those who would benefit most and not be given to those who would either not be helped or would be harmed by it. A lot of attention in precision medicine focuses on powerful machine learning algorithms that can identify new sub-populations of patient groups. However, the results depend on the input data. The input data is defined by which patients are included. Deciding which patients to include in an analysis raises important ethical issues about who is and is not being best served by innovations in the field.
There are moral, scientific and economic reasons for having a more inclusive patient demographic represented in precision medicine.
From a moral view
We want to support technologies that could benefit all of society and we want to reduce the potential harm to future patients. Including broader demographic coverage of patients means we are more likely to provide solutions that better target their needs. It also means we are better able to identify groups who would not benefit from or be harmed by a drug being developed.
From a scientific view
We know that the effectiveness of drug treatments can be influenced by a patient’s age, sex, ethnicity or other demographic factors. There are also indirect factors that are important. Race and socio-economic status lack a biological meaning, but they are linked with long-standing healthcare inequities and challenges with recruiting participants for clinical trials.
From an economic view
Having a more inclusive patient representation in precision medicine activities provides benefits on both the demand and supply sides of pharmaceuticals. On the demand side, healthcare bodies can rationalise limited financial resources to buy medicines that are most likely to work for patient groups. With wider demographic coverage in the data sources that lead to innovations, the buyers may be able to serve more people who are part of their general patient populations. On the supply side, having more diverse data sources available will make it more likely to produce treatments that will pass clinical trials and support new patient markets.
All of these reasons address why it would be good to improve the diversity of patients represented in patient data. However, it is also important to show how diversity could be improved. The current lack of diversity in the field is recognised and widely discussed in areas of policy development, research project funding, and drug approval.
The UK Bioindustry Association (BIA) recognises that the first step to improving data diversity is knowing how diverse acquired or available datasets are. This awareness and understanding enables the researcher to work to plug the data gap. To enable this assessment, BenevolentAI has provided a pragmatic solution:
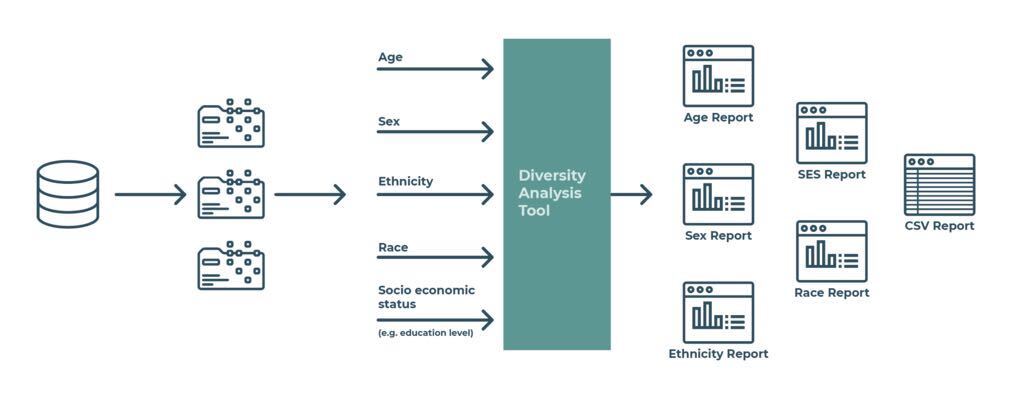
Figure 1 - outline of what the Diversity Analysis Tool does
The program works as shown in the diagram. Suppose you are in an organisation that uses patient-level data to drive a precision medicine activity. You may link various data sets that contain the age, sex, ethnicity, race and socio-economic status of each patient. In doing so, you produce a single data set that contains these variables. The program will transform the demographic data set and use it to generate multiple graphs (see figures 2 and 3). It also generates another CSV file which groups patient records by age bands. The program can be run on its own, or you can use it as a library that you include in other programs. Before you use it, you should ensure that it will support good information governance practices.
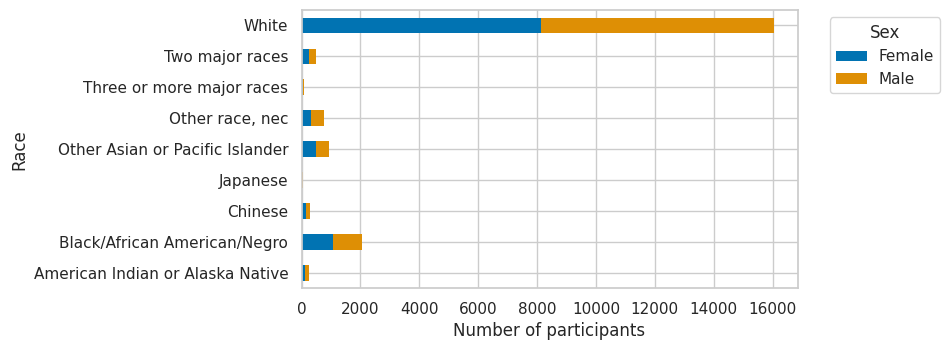
Figure 2 - an example output showing the race and sex of samples in IPUMS USA, University of Minnesota
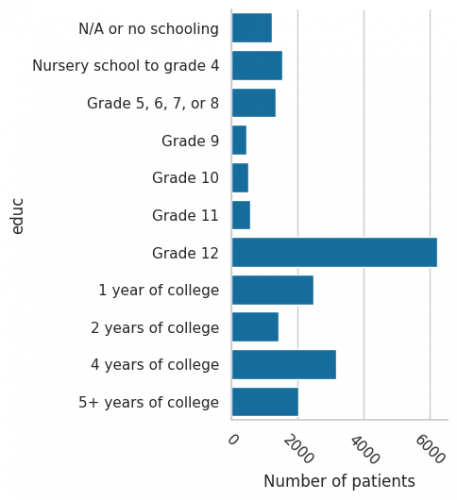
Figure 3 - an example output showing the educational attainment of samples in IPUMS USA, University of Minnesota, an indication of SES
This scenario we describe here oversimplifies some of the work that is typically needed to properly process demographic data. The program makes assumptions about the input data that may vary depending on the health data being used. It also only produces results for a single data set rather than trying to pool results from multiple data sets. Significant work is often needed to harmonise codes and categories for ethnicity, race and socio-economic status that vary among data sets.
The code base we have open-sourced is meant to be simple and provide some basic code examples that may inspire you to develop more sophisticated solutions. We also provide information about common data processing issues that relate to these demographic concepts. The code base may help you make a better assessment of the diversity in data sets you already work with. It may just help you evaluate the diversity in future health data sets you may encounter. However you may use the code, we hope it encourages you to think more about improving the diversity in data.

Open Source programme
The Diversity Analysis Tool is available on GitHub here
Brought to you by the BenevolentAI team: Mario Black, Monica Belich, Reubin Bhogal, Aylin Cakiroglu, Lucas Eggers, Matyas Fodor, Kevin Garwood, Alexander Gaujean, Gloria Hwang, Rajin Kang, Celine Lature, Nicholas Litombe, Alix Lacoste, Adepeju Oshisanya, Ibukunoluwa Oluwayomi, Aishaini Puvanendran, Anthony Risidore
Back to blog post and videos